- Medical College, Aga Khan University, Karachi, Sindh, Pakistan,
- Department of Pediatrics, Aga Khan University, Karachi, Sindh, Pakistan.
- Department of Neurosurgery, Aga Khan University, Karachi, Sindh, Pakistan.
Correspondence Address:
Kaleem Sohail Ahmed, Medical College, Aga Khan University, Karachi, Sindh, Pakistan.
DOI:10.25259/SNI_433_2021
Copyright: © 2021 Surgical Neurology International This is an open-access article distributed under the terms of the Creative Commons Attribution-Non Commercial-Share Alike 4.0 License, which allows others to remix, tweak, and build upon the work non-commercially, as long as the author is credited and the new creations are licensed under the identical terms.How to cite this article: Adnan A. Khan1, Hamza Ibad1, Kaleem Sohail Ahmed1, Zahra Hoodbhoy2, Shahzad M. Shamim3. Deep learning applications in neuro-oncology. 30-Aug-2021;12:435
How to cite this URL: Adnan A. Khan1, Hamza Ibad1, Kaleem Sohail Ahmed1, Zahra Hoodbhoy2, Shahzad M. Shamim3. Deep learning applications in neuro-oncology. 30-Aug-2021;12:435. Available from: https://surgicalneurologyint.com/surgicalint-articles/11074/
Abstract
Deep learning (DL) is a relatively newer subdomain of machine learning (ML) with incredible potential for certain applications in the medical field. Given recent advances in its use in neuro-oncology, its role in diagnosing, prognosticating, and managing the care of cancer patients has been the subject of many research studies. The gamut of studies has shown that the landscape of algorithmic methods is constantly improving with each iteration from its inception. With the increase in the availability of high-quality data, more training sets will allow for higher fidelity models. However, logistical and ethical concerns over a prospective trial comparing prognostic abilities of DL and physicians severely limit the ability of this technology to be widely adopted. One of the medical tenets is judgment, a facet of medical decision making in DL that is often missing because of its inherent nature as a “black box.” A natural distrust for newer technology, combined with a lack of autonomy that is normally expected in our current medical practices, is just one of several important limitations in implementation. In our review, we will first define and outline the different types of artificial intelligence (AI) as well as the role of AI in the current advances of clinical medicine. We briefly highlight several of the salient studies using different methods of DL in the realm of neuroradiology and summarize the key findings and challenges faced when using this nascent technology, particularly ethical challenges that could be faced by users of DL.
Keywords: Deep learning, Glioma prognostication, Machine learning, Neuro-oncology
INTRODUCTION
Amidst contemporary shifts in the global “smart-tech” arena, there has been a parallel shift in the dynamics of technological integration in health care and its rapidly growing role in supplementing traditional methods of patient care.[
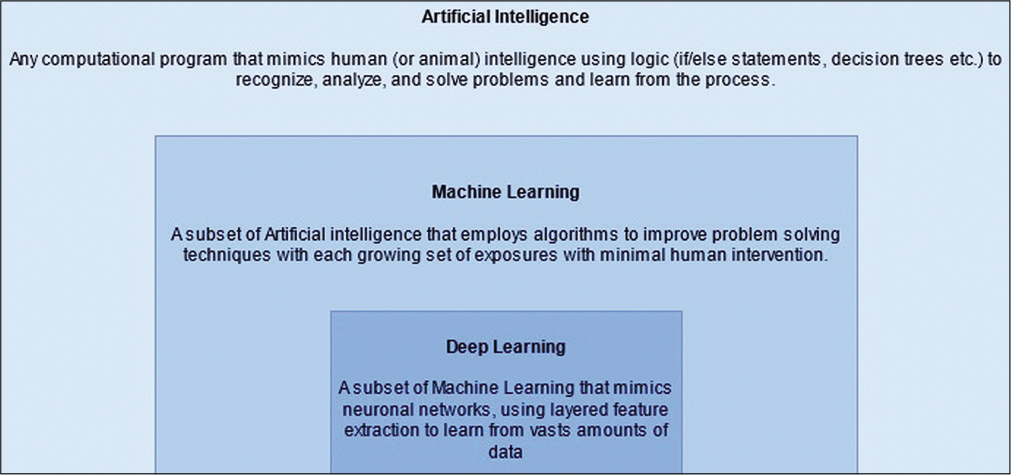
The realm of AI contains within itself machine learning (ML), as well as a further subset known as deep learning (DL). ML takes use of statistical models to enable these algorithms to improve as more data are introduced to them [
DL algorithms may then function when provided sets of parameters to evaluate a plethora of aspects of clinical importance in medicine, including screening, diagnosis, and treatment efficacy. Each algorithm must first be assessed by the relevant performance metrics of the extracted patterns when extrapolating beyond the model dataset.[
Consequently, this generalizability requires a revolution in data mining to generate large volumes of data from which this subset of AI technology can learn. This is vital to overcome the challenges of data sparsity, multicollinearity, and overfitting, and therefore, to effectively minimize internal bias in the model developed.[
Based on a subtle balance between use and limitations, DL promises to play a crucial role in healthcare, allowing us to find a niche between generalized practice guidelines and personalized patient-centric care. These algorithms have already shown a high level of diagnostic performance based on imaging data for conditions including diabetic retinopathy, skin cancer, and pneumonia.[
Amidst these enhancements in health care across disciplines, the realm of prediction and management in oncology stands to benefit greatly from such technologies.[
DIAGNOSIS, GRADING, AND CLASSIFICATION OF GLIOMAS
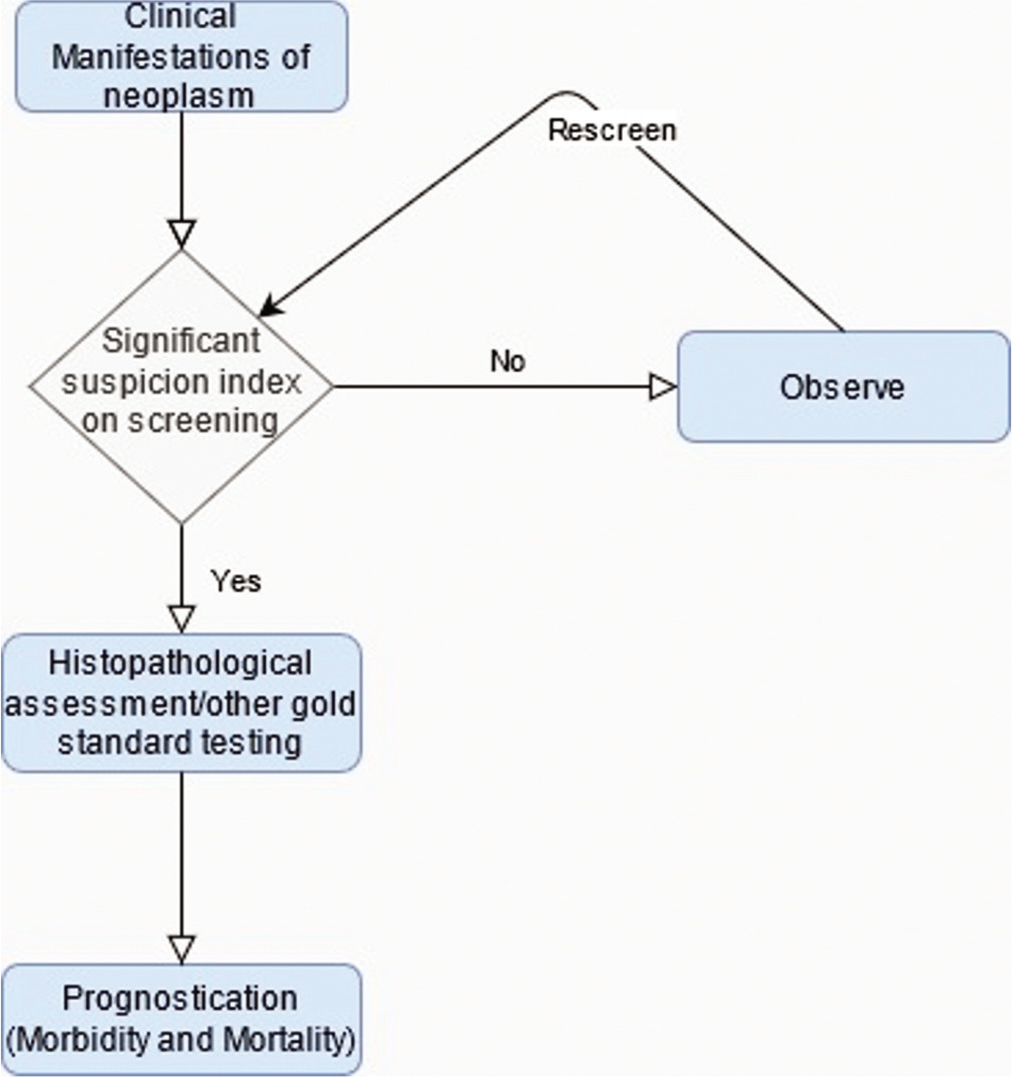
The clinical course of a neoplasm is dependent on the efficacy of several stages of medical intervention including the recognition of clinical manifestations, initial screening of the lesion (employing imaging or laboratory testing), the decision to observe or treat, postscreening histopathological assessment/gold standard confirmation, and predicting morbidity and mortality outcomes. ML can be used to aid this paradigm of medical intervention at several of these levels [
As mentioned before, the strength of ML as a tool to evaluate radiological screening independently or augmentatively is yielding promising results, with this excitement not only limited to mammography but also in the realms of lung cancer, prostate cancer, and brain metastases with the use of CADe and computer-aided diagnoses techniques.[
The promise of ML in the field of oncology in addition to the growing recognition of an integrated approach to neuropathology lends optimism to the treatment of a subset of neuro-oncological entities known as gliomas. These tumors are difficult to render an exact judgment on, due to similarity of cellular structures among lower grade lesions; with estimations varying due to inter-reader variability depending on the professional experience of histopathologists and radiologists, introducing an element of subjectivity.[
Broadly, ML techniques can be divided into those that learn to predict outcomes based on input-output paired training, termed supervised learning, or those that find patterns within input data itself without preset outcomes, termed unsupervised learning. Supervised learning itself can be achieved through several techniques such as support vector matrixes (SVMs) which aim to delineate a function that separates two sets of data.[
Previously, studies have demonstrated that gliomas can be classified according to their clinical grade using linear SVMs which were trained on descriptive features such as amount of mass effect of blood supply. The limitation of these studies, such as one by Li et al., is that these quantitative features were estimated by domain experts and the definition of these features was limited to expert opinion and hence not reproducible.[
It stands to reason from this discussion that the beginnings of ML in neuropathology must be the generation of a dataset from which newly developed AI can learn. For gliomas, radiological evidence in addition to matched clinical, genetic, and pathological data collections reside in the Cancer Imaging Archive and Genomics Data Commons Data Portal respectively, as a joint effort between the National Cancer Institute and the National Human Genome Research Institute from 2006 onward.[
At present, algorithms generated by ML have shown promise in accurate recognition of characteristics used to define gliomas. Noninvasive neuroimaging tools for glioma grading using a multitude of quantitative parameters obtained from advanced magnetic resonance imaging (MRI) techniques, such as dynamic contrast-enhanced MRI, arterial spin labeling, and diffusion-weighted imaging, are currently being utilized in many ML/DL models.[
The emerging field of radiogenomics breaks the restrictions of applying AI technology to segmentation only. Advancement in our understanding of key molecular differences in tumor cells in comparison to normal cells and how these differences drive changes in the microenvironment of the lesions now lends itself to radiological patterns that can more accurately identify a tumor. For example, certain glioma varieties exhibit differences in isocitrate dehydrogenase (IDH), a key enzyme in the Krebs cycle. Gliomas that potentiate the activity of certain genotypes can drive the production of 2-hydroxyglutaric acid, vascular endothelial growth factor, and hypoxia-induced factor-1a. The activity of these molecules may, in turn, produce characteristic radiological findings such as alterations in histogram analysis.[
In fact, the current literature supports the idea of using radiological imaging to predict the presence of such biomolecular differences. Specifically in the context of gliomas, studies have been conducted on IDH mutations, O6-methylguanine methyltransferase (MGMT) hypermethylation, epidermal growth factor receptors (EGFRs), and 1p/19q chromosomal codeletions.
Zhang et al. were able to generate a predictive model for IDH genotypes (of which genotypes 1 and 2 predict good response to therapy) within high-grade glioma lesions based on clinical data and radiological features obtained through conventional MRI with an AUROC of 0.9231 in a validation cohort.[
Similarly to IDH, MGMT hypermethylation is an important identifying marker due that also happens to predict higher response to treatment with combination temozolomide and radiation.[
Other studies have also shown the ability of ML to detect 1p19q chromosomal codeletion (recognized as predictor of chemotherapeutic response in gliomas), and EGFR amplification status, another avenue for targeted therapy. The aforementioned neural network developed by Chang et al. manages to predict 1p19q codeletion status at an accuracy of 92%.[
With such exciting advancements, comparable outputs to traditional methods of algorithmic generation, and a yet unmet ceiling for improvement, it is thought that in the near future, with the increasing availability of high-quality data, CNNs will be the mainstay in both diagnosing and prognosticating cancer, using widely available imaging modalities.
PROGNOSIS PREDICTION
While ML can be useful in predicting prognosis based just on imaging characteristics before surgery, we have more information after surgical resection of the tumor that can be incorporated in making extensive ML algorithms. Along with imaging characteristics, the cumulative biopsy and clinical data such as tumor markers can help generate ML algorithms for estimating prognosis, complications, and other outcomes. It can also overcome challenges such as the correct recognition of tumor progression versus pseudoprogression, a transient MRI finding mimicking progression with eventual improvement. One such study aiming to delineate the two conducted by Jang et al. developed a CNN model with an AUROC of 83% that was trained on 59 individuals and tested on 19 (Seoul National University Hospital dataset).[
Conventionally, prognosticating for GBM involves accounting for a multitude of independent risk factors impacting overall survival (OS) such as male gender, age >60 years), poor preoperative Karnofsky scores of <70, Caucasian ethnicity, advanced tumor with partial resection, and surgery without adjuvant chemoradiation.[
A study by Nie et al. demonstrated that through a 3D CNN, they were able to train a support vector machine to predict a long or short OS time. Their experimental results were able to achieve 89.9% accuracy through their methods.[
Another study by Zhou et al. utilized nonquantitative spatial-correlated features from MRI defined tumor subregions (termed habitats) in developing a computational framework. This framework was able to use intratumoral grouping and spatial mapping to identify GBM tumor subregions and yield habitat-based features. After separating data sets into those that underwent resection with GBM (32 patients), and those that did not (22 patients), they were able to achieve 87.50% and 86.36% accuracies for survival group prediction, respectively.[
The role of computational networks in prognosticating overall long- and short-term survival is one that has been evaluated through dozens of studies in low- to mid-grade tumors, yielding similar results when applied to high-grade tumors as well.[
CHALLENGES AND LIMITATIONS
Ethical considerations
ML, like any early adopted technology, is not without its potential drawbacks in terms of ethical dilemmas.[
There is an incredibly urgent need for ethical guidelines for medical practitioners to use these advancing AI and machine learning technologies. Time-constrained consultants are expected to understand the inner workings of these insular programs, as without knowing the data sets they are built on, the technology can progress into a black box leading to ethically gray outcomes. The lack of transparency inherent with these systems may lead to physicians doubt either the algorithm, or themselves, when their decision is conflicting with one made by a machine.
It is also important to recall that the foundation of the current medical system in many nations was based on a patient-doctor relationship, not a patient-health-care system relationship. Many doctors (and patients) feel uneasy at the growing rate at which we are adopting these machines into our health-care decisions. One study showed that over 63% of adults in the United Kingdom felt uncomfortable with AI replacing the conventional decision-making process.[
Finally, the laws and rules of ethics are a worldwide debate but enforced by national laws. Ethical laws on the potential of AI and ML in medicine have been hotly contentious but are virtually absent in low- and middle-income countries.[
Another reason for the impediment in widespread adoption, and perhaps the most important one, is a lack of accountability.[
Other logistical drawbacks to the use of AI in medicine include the scale of computational power required to develop useful algorithms and prioritization of resource allocation toward more pragmatic algorithms to maximize their effects.
Specifically in the field of radiological AI interpretations, a major drawback in considering findings and pattern recognition in any image modality is a lack of consensus on which features would add most value to a preoperative diagnosis, whether from a histogram parameters or image texture attributes.[
A prospective trial on glioma patients, using a previously trained CNN based on an accurate model, to help prognosticate or diagnose a patient is sorely needed. Until, we overcome challenges in implementation and their exact use in clinical decision-making processes becomes apparent, they will remain promising research ventures.
CONCLUSION
The use of AI to augment the clinical judgment and practice of a physician is an exciting prospect, specifically in the field of neuro-oncology where neuroradiology and neuroradiogenomics are prime candidates for the use of the algorithmic learning in order. In a noninvasive manner, these tools can predict the presence of several factors that can be helpful in the diagnosis and prognostication of gliomas. Current literature already demonstrates a high sensitivity and accuracy of individual learning techniques in determining genetic markers and radiological features. The challenge lies in creating algorithms that are applicable on much larger scale, with greater amounts of learning and practice sets for AI learning techniques to be tested on, while still remaining logistically feasible to be run within the strict timeframes within which health-care systems operate. In addition, there are ethical considerations to be made regarding the use of AI within the clinical realm and their impact on a physician’s decision-making.
Declaration of patient consent
Patient’s consent not required as there are no patients in this study.
Financial support and sponsorship
Nil.
Conflicts of interest
There are no conflicts of interest.
References
1. Akkus Z, Ali I, Sedlář J, Agrawal JP, Parney IF, Giannini C. Predicting deletion of chromosomal arms 1p/19q in low-grade gliomas from MR images using machine intelligence. J Digit Imaging. 2017. 30: 469-76
2. Ambrosini RD, Wang P, O’Dell WG. Computer-aided detection of metastatic brain tumors using automated three-dimensional template matching. J Magn Reson Imaging. 2010. 31: 85-93
3. Baştanlar Y, Özuysal M. Introduction to machine learning. Methods Mol Biol. 2014. 1107: 105-28
4. Bi WL, Hosny A, Schabath MB, Giger ML, Birkbak NJ, Mehrtash A. Artificial intelligence in cancer imaging: Clinical challenges and applications. CA Cancer J Clin. 2019. 69: 127-57
5. Bzdok D, Krzywinski M, Altman N. Points of significance: Machine learning: A primer. Nat Methods. 2017. 14: 1119-20
6. Caruana R, Caruana R, Freitag D.editors. Greedy attribute selection. Proceedings of the elevator International Conference Machine Learning. 1994. p. 28-36
7. Chan HP, Hadjiiski L, Zhou C, Sahiner B. Computer-aided diagnosis of lung cancer and pulmonary embolism in computed tomography-a review. Acad Radiol. 2008. 15: 535-55
8. Chang K, Bai HX, Zhou H, Su C, Bi WL, Agbodza E. Residual convolutional neural network for the determination of IDH status in low-and high-grade gliomas from mr imaging. Clin Cancer Res. 2018. 24: 1073-81
9. Chang P, Grinband J, Weinberg BD, Bardis M, Khy M, Cadena G. Deep-learning convolutional neural networks accurately classify genetic mutations in gliomas. AJNR Am J Neuroradiol. 2018. 39: 1201-7
10. Char DS, Shah NH, Magnus D. Implementing machine learning in health care-addressing ethical challenges. N Engl J Med. 2018. 378: 981-3
11. Devos A, Simonetti AW, van der Graaf M, Lukas L, Suykens JA, Vanhamme L. The use of multivariate MR imaging intensities versus metabolic data from MR spectroscopic imaging for brain tumour classification. J Magn Reson. 2005. 173: 218-28
12. Fenech M, Strukelj N, Buston O.editors. Ethical Social and Political Challenges of Artificial Intelligence in Health. 2018. p.
13. Freymann J. TCGA Glioma Phenotype Research Group-The Cancer Imaging Archive (TCIA) Public Access-Cancer Imaging Archive Wiki. Available from: https://www.wiki.cancerimagingarchive.net/display/public/tcga+glioma+phenotype+research+group [Last accessed on 2021 Apr 25].
14. Gittleman H, Cioffi G, Chunduru P, Molinaro AM, Berger MS, Sloan AE. An independently validated nomogram for isocitrate dehydrogenase-wild-type glioblastoma patient survival. Neurooncol Adv. 2019. 1: vdz007
15. Guo J, Li B. The application of medical artificial intelligence technology in rural areas of developing countries. Health Equity. 2018. 2: 174-81
16. Han L, Kamdar MR. MRI to MGMT: Predicting methylation status in glioblastoma patients using convolutional recurrent neural networks. Pac Symp Biocomput. 2018. 23: 331-42
17. Hegi ME, Diserens AC, Gorlia T, Hamou MF, de Tribolet N, Weller M. MGMT gene silencing and benefit from temozolomide in glioblastoma. N Engl J Med. 2005. 352: 997-1003
18. Hoodbhoy Z, Hasan B, Siddiqui K. Does artificial intelligence have any role in healthcare in low resource settings?. J Med Artif Intell. 2019. 2: 13
19. Jang BS, Jeon SH, Kim IH, Kim IA. Prediction of pseudoprogression versus progression using machine learning algorithm in glioblastoma. Sci Rep. 2018. 8: 12516
20. Jang BS, Park AJ, Jeon SH, Kim IH, Lim DH, Park SH. Machine learning model to predict pseudoprogression versus progression in glioblastoma using mri: A multi-institutional study (KROG 18-07). Cancers (Basel). 2020. 12: 2706
21. Kickingereder P, Bonekamp D, Nowosielski M, Kratz A, Sill M, Burth S. Radiogenomics of glioblastoma: Machine learning-based classification of molecular characteristics by using multiparametric and multiregional MR imaging features. Radiology. 2016. 281: 907-18
22. Korfiatis P, Kline TL, Lachance DH, Parney IF, Buckner JC, Erickson BJ. Residual deep convolutional neural network predicts MGMT methylation status. J Digit Imaging. 2017. 30: 622-8
23. Li GZ, Yang J, Ye CZ, Geng DY. Degree prediction of malignancy in brain glioma using support vector machines. Comput Biol Med. 2006. 36: 313-25
24. Louis DN, Perry A, Guido R, von Deimling A, FigarellaBranger D, Webster K. The 2016 World Health Organization classification of tumors of the central nervous system: A summary. Acta Neuropathol. 2016. 131: 803-20
25. Machine Bias-ProPublica. Available from: https://www.propublica.org/article/machine-bias-risk-assessments-incriminal-sentencing [Last accessed on 2021 Apr 25].
26. Ngiam KY, Khor IW. Big data and machine learning algorithms for health-care delivery. Lancet Oncol. 2019. 20: e262-73
27. Nie D, Zhang H, Adeli E, Liu L, Shen D.editors. 3D deep learning for multi-modal imaging-guided survival time prediction of brain tumor patients, Lecture Notes in Computer Science (Including Subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics). Berlin, Germany: Springer; 2016. 9901: 212-20
28. Panch T, Szolovits P, Atun R. Artificial intelligence, machine learning and health systems. J Glob Health. 2018. 8: 020303
29. Rajendran P, Madheswaran M. An improved image mining technique for brain tumour classification using efficient classifier. Int J Comput Sci Inf Secur. 2010. 6: 1-10
30. Salim M, Wåhlin E, Dembrower K, Azavedo E, Foukakis T, Liu Y. External evaluation of 3 commercial artificial intelligence algorithms for independent assessment of screening mammograms. JAMA Oncol. 2020. 6: 1581-8
31. Sanghani P, Ang BT, King NK, Ren H. Overall survival prediction in glioblastoma multiforme patients from volumetric, shape and texture features using machine learning. Surg Oncol. 2018. 27: 709-14
32. Skogen K, Schulz A, Dormagen JB, Ganeshan B, Helseth E, Server A. Diagnostic performance of texture analysis on MRI in grading cerebral gliomas. Eur J Radiol. 2016. 85: 824-9
33. Sotoudeh H, Shafaat O, Bernstock JD, Brooks MD, Elsayed GA, Chen JA. Artificial intelligence in the management of glioma: Era of personalized medicine. Front Oncol. 2019. 9: 768
34. Valdebenito J, Medina F. Machine learning approaches to study glioblastoma: A review of the last decade of applications. Cancer Rep. 2019. 2: e1226
35. Wang J, Hu G, Quan X. Analysis of the factors affecting the prognosis of glioma patients. Open Med. 2019. 14: 331-5
36. Watson DS, Krutzinna J, Bruce IN, Griffiths CE, McInnes IB, Barnes MR. Clinical applications of machine learning algorithms: Beyond the black box. BMJ. 2019. 364: l886
37. Whittle IR. The dilemma of low grade glioma. J Neurol Neurosurg Psychiatry. 2004. 75: ii31-6
38. Xu C, Jackson SA. Machine learning and complex biological data. Genome Biol. 2019. 20: 76
39. Yang S, Zhu F, Ling X, Liu Q, Zhao P. Intelligent health care: Applications of deep learning in computational medicine. Front Genet. 2021. 12: 607471
40. Zacharaki EI, Kanas VG, Davatzikos C. Investigating machine learning techniques for MRI-based classification of brain neoplasms. Int J Comput Assist Radiol Surg. 2011. 6: 821-8
41. Zhang B, Chang K, Ramkissoon S, Tanguturi S, Bi WL, Reardon DA. Multimodal MRI features predict isocitrate dehydrogenase genotype in high-grade gliomas. Neuro Oncol. 2017. 19: 109-17
42. Zhou H, Vallières M, Bai HX, Su C, Tang H, Oldridge D. MRI features predict survival and molecular markers in diffuse lower-grade gliomas. Neuro Oncol. 2017. 19: 862-70
43. Zhou M, Scott J, Chaudhury B, Hall L, Goldgof D, Yeom KW. Radiomics in brain tumor: Image assessment, quantitative feature descriptors, and machine-learning approaches. Am J Neuroradiol. 2018. 39: 208-16
44. Zhou M, Chaudhury B, Hall LO, Goldgof DB, Gillies RJ, Gatenby RA. Identifying spatial imaging biomarkers of glioblastoma multiforme for survival group prediction. J Magn Reson Imaging. 2017. 46: 115-23