- Department of Neurosurgery, University of New Mexico, Albuquerque, New Mexico, United States
- Department of Neurosurgery, School of Medicine, New York Medical College, Valhalla, New York, United States,
- Department of Neurosurgery, Topiwala National Medical and B. Y. L. Nair Charitable Hospital, Mumbai, Maharashtra, India.
Correspondence Address:
Evan Courville, Department of Neurosurgery, University of New Mexico, Albuquerque, New Mexico, United States.
DOI:10.25259/SNI_312_2023
Copyright: © 2023 Surgical Neurology International This is an open-access article distributed under the terms of the Creative Commons Attribution-Non Commercial-Share Alike 4.0 License, which allows others to remix, transform, and build upon the work non-commercially, as long as the author is credited and the new creations are licensed under the identical terms.How to cite this article: Evan Courville1, Syed Faraz Kazim1, John Vellek2, Omar Tarawneh2, Julia Stack2, Katie Roster2, Joanna Roy3, Meic Schmidt1, Christian Bowers1. Machine learning algorithms for predicting outcomes of traumatic brain injury: A systematic review and meta-analysis. 28-Jul-2023;14:262
How to cite this URL: Evan Courville1, Syed Faraz Kazim1, John Vellek2, Omar Tarawneh2, Julia Stack2, Katie Roster2, Joanna Roy3, Meic Schmidt1, Christian Bowers1. Machine learning algorithms for predicting outcomes of traumatic brain injury: A systematic review and meta-analysis. 28-Jul-2023;14:262. Available from: https://surgicalneurologyint.com/surgicalint-articles/12468/
Abstract
Background: Traumatic brain injury (TBI) is a leading cause of death and disability worldwide. The use of machine learning (ML) has emerged as a key advancement in TBI management. This study aimed to identify ML models with demonstrated effectiveness in predicting TBI outcomes.
Methods: We conducted a systematic review in accordance with the Preferred Reporting Items for Systematic Review and Meta-Analysis statement. In total, 15 articles were identified using the search strategy. Patient demographics, clinical status, ML outcome variables, and predictive characteristics were extracted. A small meta-analysis of mortality prediction was performed, and a meta-analysis of diagnostic accuracy was conducted for ML algorithms used across multiple studies.
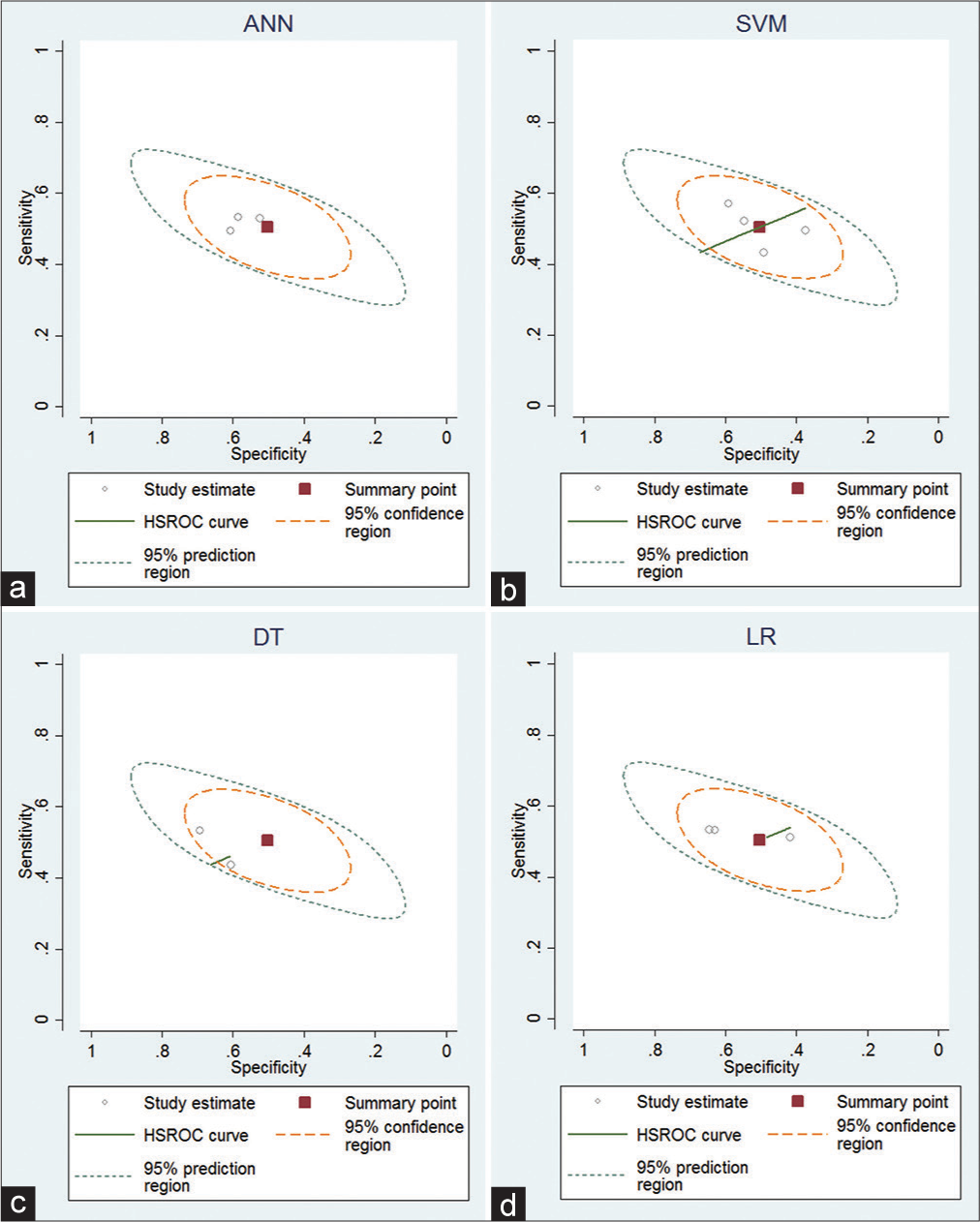
Results: ML algorithms including support vector machine (SVM), artificial neural networks (ANN), random forest, and Naïve Bayes were compared to logistic regression (LR). Thirteen studies found significant improvement in prognostic capability using ML versus LR. The accuracy of the above algorithms was consistently over 80% when predicting mortality and unfavorable outcome measured by Glasgow Outcome Scale. Receiver operating characteristic curves analyzing the sensitivity of ANN, SVM, decision tree, and LR demonstrated consistent findings across studies. Lower admission Glasgow Coma Scale (GCS), older age, elevated serum acid, and abnormal glucose were associated with increased adverse outcomes and had the most significant impact on ML algorithms.
Conclusion: ML algorithms were stronger than traditional regression models in predicting adverse outcomes. Admission GCS, age, and serum metabolites all have strong predictive power when used with ML and should be considered important components of TBI risk stratification.
Keywords: Artificial intelligence, Head injury, Machine learning, Mortality, Outcomes, Traumatic brain injury
INTRODUCTION
Physicians are often presented with large quantities of complex data and limited processing time. This presents barriers to the real-time analysis and prediction of patient outcomes. In computer science, complex algorithms designed to learn from data and create generalizations are known as machine learning (ML). The marked proliferation of electronic medical record systems during recent years has presented unique opportunities for ML to improve patient care. Several ML learning techniques have been used in clinical practice to predict deleterious events and alert appropriate care teams. This has led to an increase in the number of early interventions, reduced mortality, and decreased lengths of hospital stay.[
Traumatic brain injury (TBI) remains one most prevalent causes of death and disability throughout the world.[
MATERIALS AND METHODS
The present systematic review and meta-analysis were performed per the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) guidelines.[
Literature search strategy
We conducted a literature search of studies reporting on ML-based prediction of TBI outcomes published until March 31, 2021. We searched the following three electronic bibliographic databases: PubMed, EMBASE, and Cochrane Library. We used the following MeSH (Medical Subject Heading) terms in combination with Boolean Operators OR and AND: “machine learning” OR “artificial intelligence” OR “neural network” OR “naive Bayes” OR “Bayesian learning” OR “random forest” OR “deep learning” OR “machine intelligence” OR “boosting” OR “nature language processing” OR “decision tree” AND “traumatic brain injury” OR “head injury.” An additional search involving the following terms was also performed: “machine learning” OR “artificial intelligence” OR “neural network” OR “naive Bayes” OR “Bayesian learning” OR “random forest” OR “deep learning” OR “machine intelligence” OR “boosting” OR “nature language processing” OR “decision tree” AND “traumatic brain injury” OR “head injury” OR AND “outcome” OR “mortality” OR “morbidity.”
Inclusion and exclusion criteria
We included peer-reviewed prospective and retrospective cohort studies published in the English language utilizing ML algorithms to predict outcomes of TBI in human patients. Single case reports, editorials, reviews, and conference/meeting abstracts were excluded from the study. Furthermore, TBI studies that used ML for a purpose other than predicting outcomes were also excluded. We also reviewed the reference lists of the selected articles for any additional articles related to the topic.
Data extraction
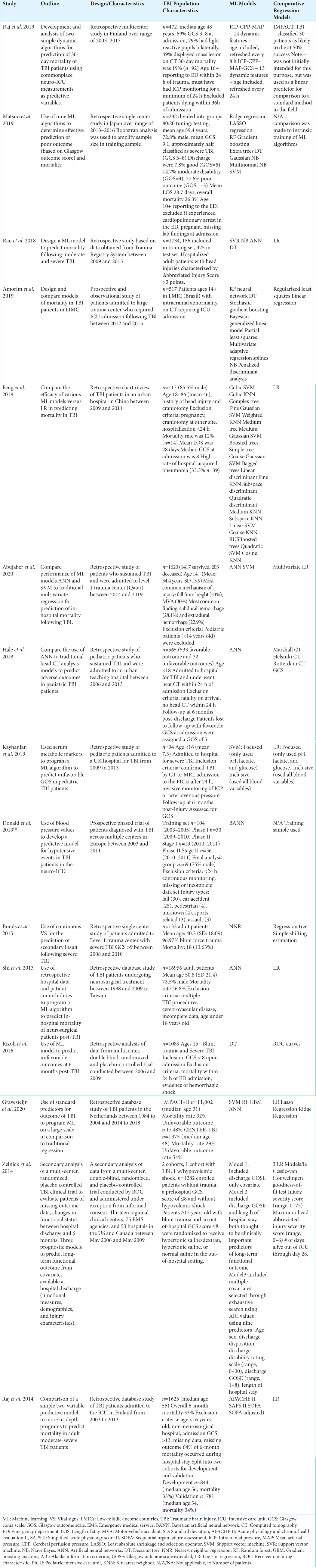
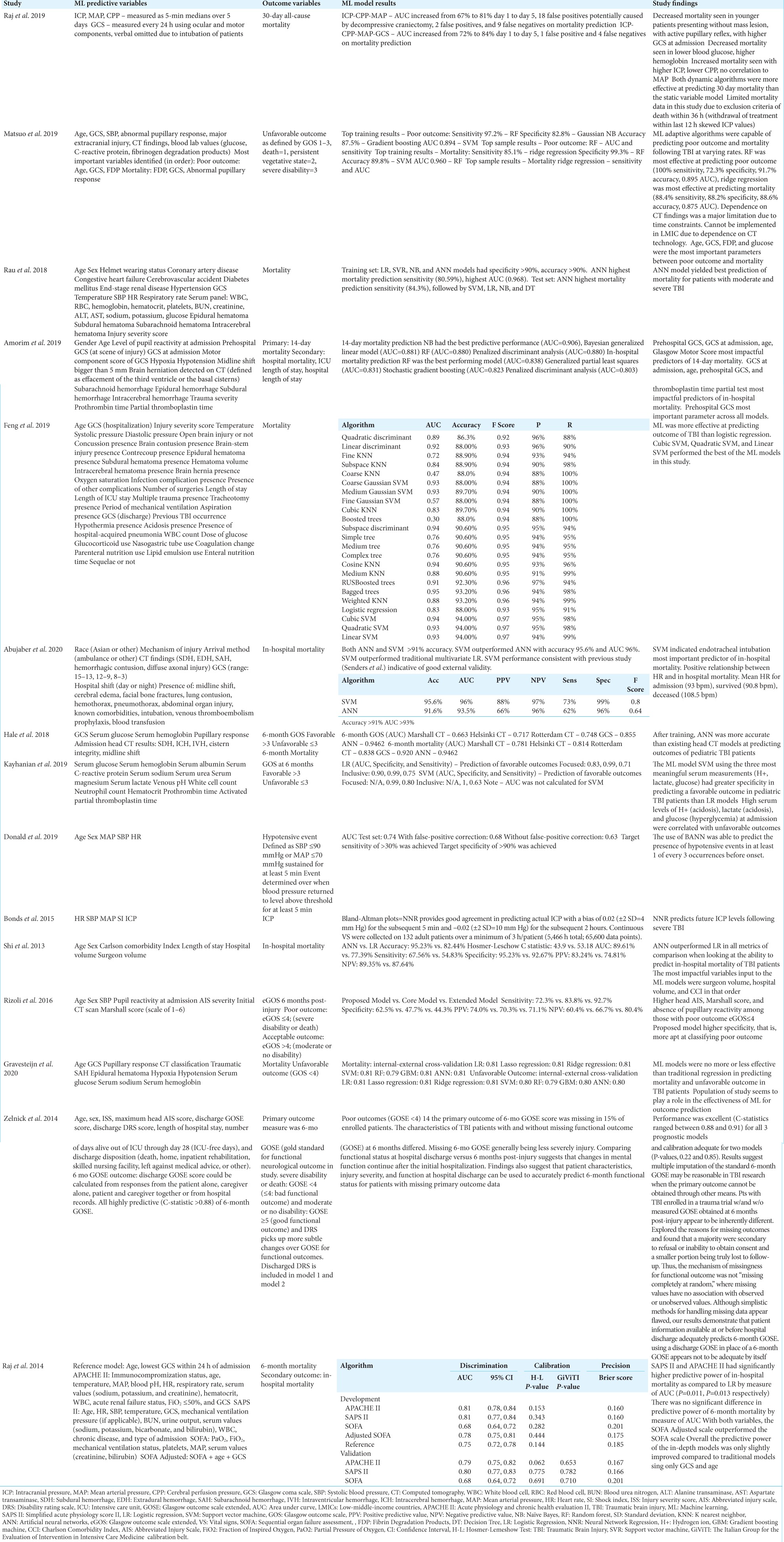
Three independent investigators (JV., OHT., and JS.) reviewed the full text of the included articles and extracted the data on a data collection form. Any disagreement between the three authors was resolved by discussion. The following data were extracted from each study: study design, TBI population characteristics, ML and comparative regression models used, ML input variables, outcome variables, study results, and predictive performance of various models used in the study.
Risk of bias assessment
We employed the Quality Assessment of Diagnostic Accuracy Studies 2 (QUADAS-2) in the Review Manager (RevMan) software version 5.4 to assess the quality of extracted studies [
Quality assessment
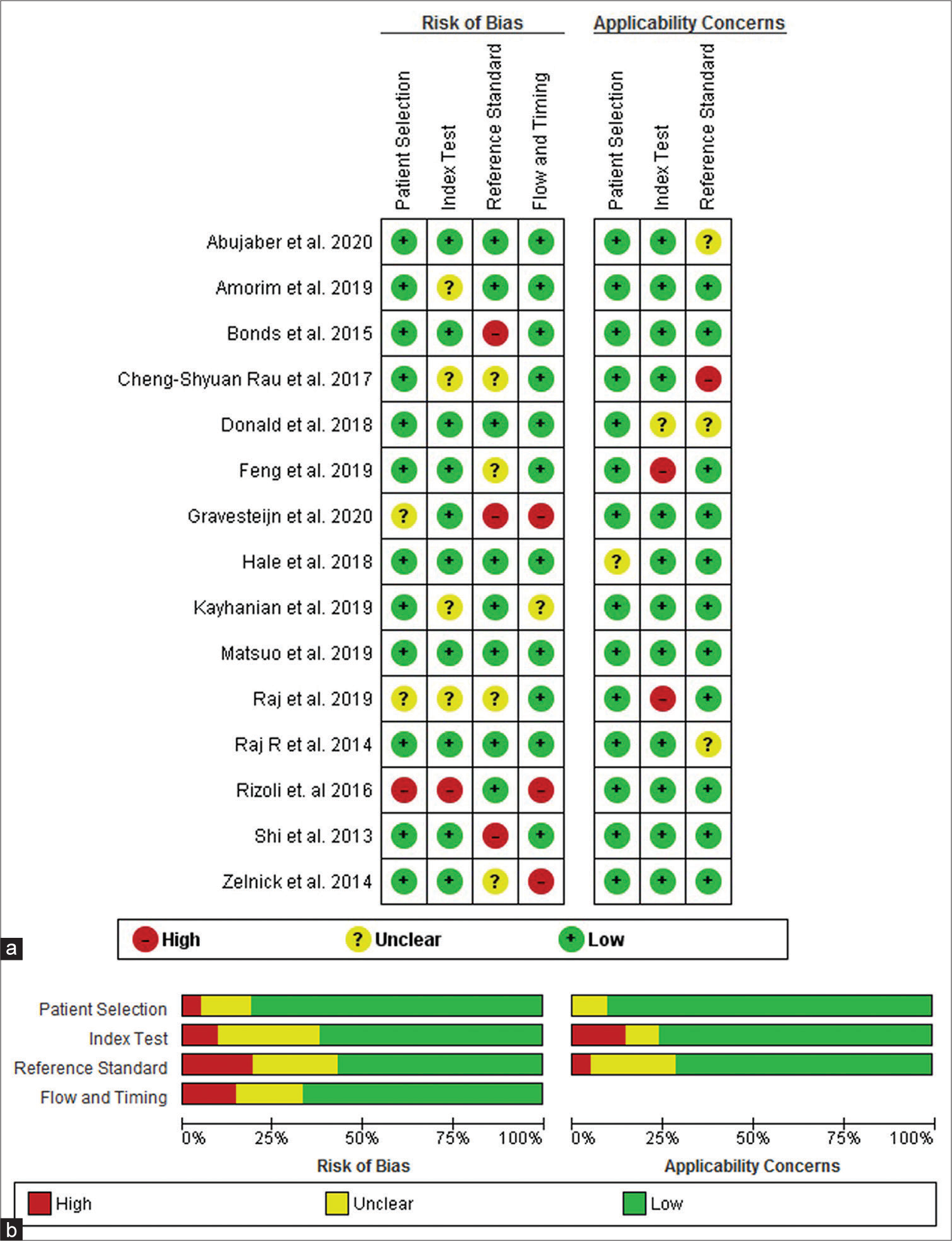
The quality of the included studies was evaluated using the QUADAS-2 tool in RevMan version 5.4.1 software. Each study was assessed using 12 signaling questions (three from each domain) and three questions regarding study applicability (one each from the first three domains) [
The domain “Patient Selection” addresses the following question: “Could the selection of patients or study participants have introduced bias?” The constitution of the study population is centrally important to a high-quality study. We distinguished three populations, study, source, and target. The study population is the population that was reported on in an article, sampled from a larger source population. Only two studies (Gravesteijn et al. 2020 and Raj et al. 2019) reported an unclear risk of bias while others answered a low risk of bias and only one study answered a high risk (Rizoli et al. 2016).[
The domain “Index Text” addresses the question: “Could the conduct or interpretation of the index test have introduced bias?” The index test results are one central component of a 2 × 2 table that is evaluated in diagnostic studies. The index test is the assay under investigation in the study, and a study may evaluate one or more index tests in the same population or among population subsets. Among the studies cohort in our systematic review and meta-analysis using ML and comparative regression models in the prediction of TBI, only one study qualified as high risk (Rizoli et al. 2016), four studies (Amorim et al. 2019, Rau et al. 2017, Kayhanian et al. 2019, and Raj et al. 2019) remained unclear risk while others reported a low risk of bias.[
The domain “Reference standard” addresses the question: “Could the reference standard, its conduct, or its interpretation have introduced bias?” Among all pooled studies, Gravesteijn et al. 2020, Shi et al. 2013, and Bonds et al. 2015 answered a high risk, Zelnick et al. 2014, Rau et al. 2018, Raj et al. 2019, and Feng et al. 2019 reported unclear risk while others pooled studies reported low risk.[
The domain “Flow and Timing” addresses the question: “Could the study flow and timing have introduced bias?” The methods and results sections should provide a clear description of clinical referral algorithms (i.e., patients who did/did not receive the index tests or reference standard, respectively) and of any patients excluded from the analyses. Three studies in our analysis (Zelnick et al. 2014, Gravesteijn et al. 2020, and Rizoli et al. 2016) reported a considerable risk of bias while one study (Kayhanian et al. 2019) answered unclear risk of bias, all others reported a low risk of bias.
Statistical analysis
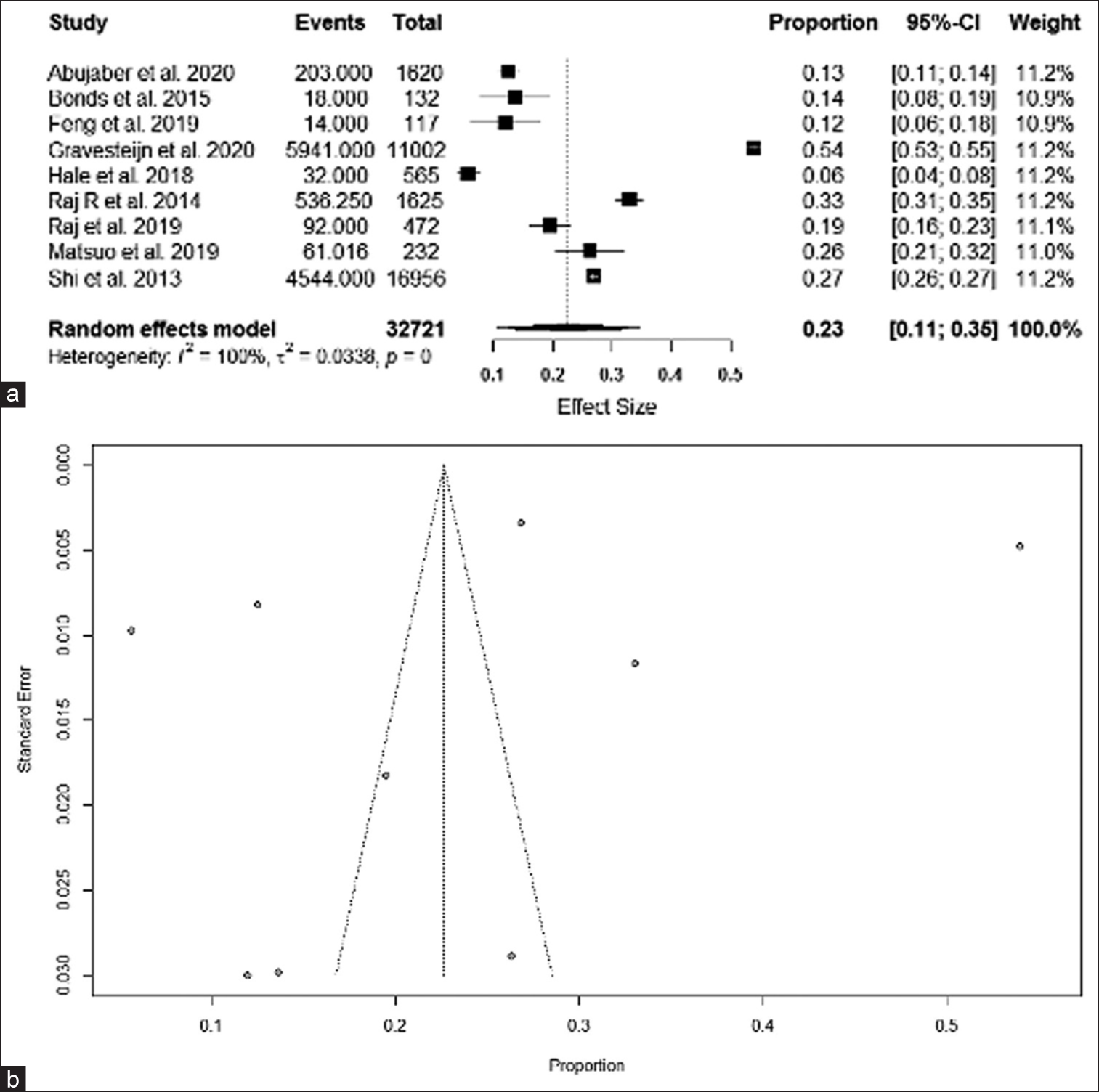
We recorded data from the included studies in a Microsoft Excel datasheet (Microsoft Corp., Redmond, Washington, USA). For the pooled mortality rate, we employed a random-effects meta-analysis model in R statistical software version 4.02 (R Foundation for Statistical Computing, Vienna, Austria). We measured the heterogeneity between the included studies employing the Higgins I2 statistic. We used a random-effects model due to the high statistical heterogeneity (defined as I2 > 25%) among studies included in the meta-analysis. Forest plots were generated using the function “metaforest” in R statistical software.[
RESULTS
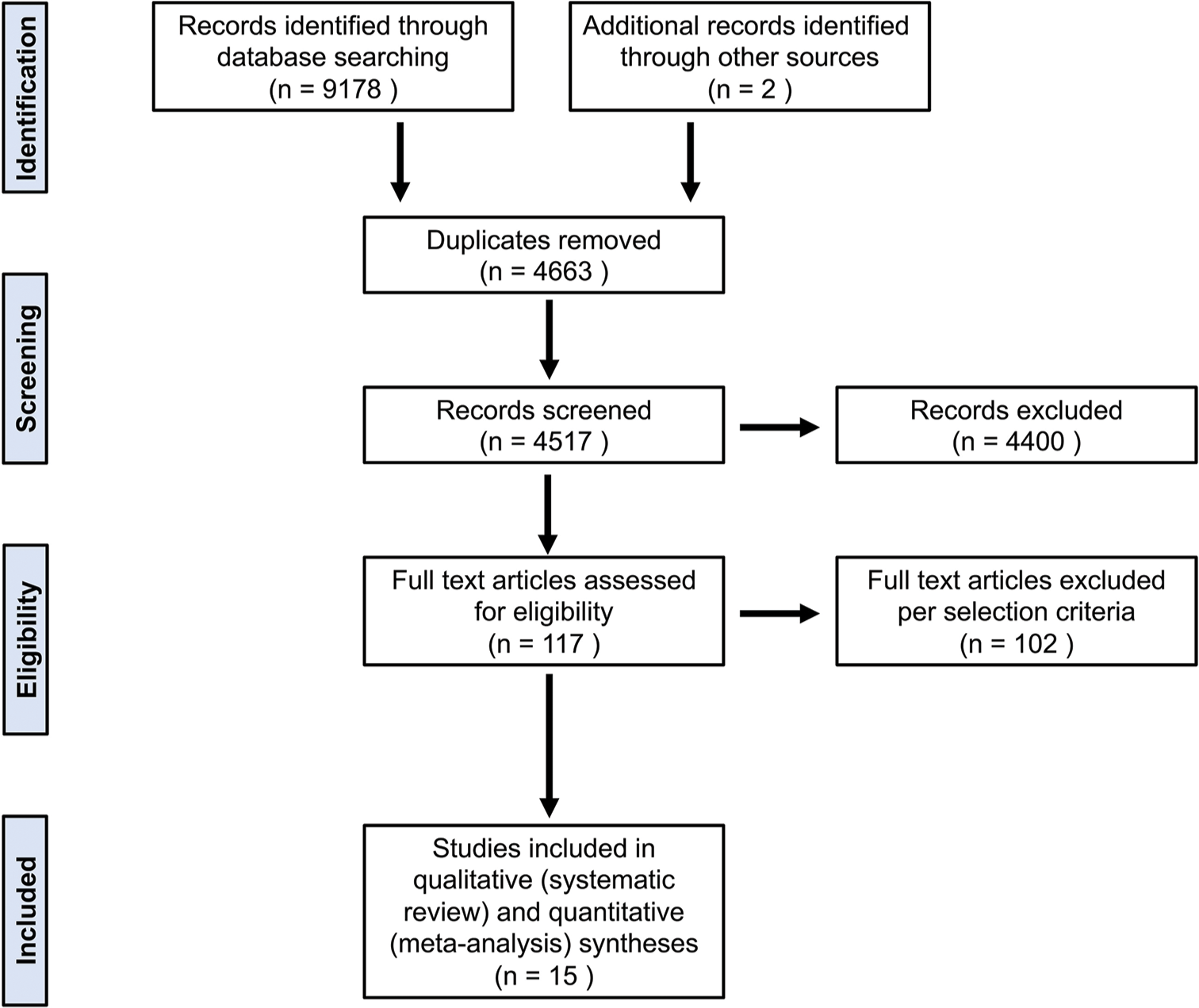
The initial literature search identified 9180 articles. After removing duplicates (n = 4663) and screening the titles and abstracts (n = 4517), we excluded a total of 9063 studies. After the screening of full-text articles based on our selection criteria (n = 117), we included a total of 15 studies in the qualitative systematic review and quantitative meta-analysis. However, the actual number of included studies for the small meta-analysis varied depending on how many studies documented the data for a particular algorithm using a similar methodology.
Prognostic factors for mortality and unfavorable outcomes
Although there was significant heterogeneity in the selected input variables used for the prediction of mortality and unfavorable outcomes, critical clinicopathological and imaging findings were identified from our review: abnormal serum glucose,[
Diagnostic accuracy: Meta-analysis for ML algorithms
A small meta-analysis was conducted for studies using mortality as a primary outcome.
In-hospital mortality
ANN and SVM have both been used to assess in-hospital mortality in a study containing 1620 patients.[
14-day mortality after TBI
An additional study used models to predict in-hospital and 14-day mortality. The models assessed mortality in 517 TBI patients in a low-middle-income country (LMIC).[
30-day mortality after TBI
Two custom models were developed to predict the 30-day mortality of TBI patients using commonplace neurointensive care unit measurements as predictive variables.[
Unfavorable outcomes at 6 months
DT was used as the ML model to predict unfavorable outcomes at 6 months post-TBI.[
Prediction of outcomes in the pediatric population
ANN was compared to traditional head computed tomography (CT) analysis (i.e., Marshall CT, Helsinki CT, and Rotterdam CT) and GCS to predict adverse outcomes and mortality in pediatric TBI patients.[
Prediction of secondary insults: ICP, hypotensive events, and shock index (SI)
Two studies used vital signs as outcome variables for ML. Bayesian Artificial Neural Network (BANN) was used to assess blood pressure values to develop a predictive model for hypotensive events in TBI patients in the neuro-intensive care unit.[
DISCUSSION
In the present systematic review and meta-analysis, we evaluate the predictive power of various ML algorithms for unfavorable outcomes and mortality in patients with TBI. Several studies have demonstrated the utility of ML in medicine; however, most TBI studies were focused on diagnosis and classification.[
TBI remains one of the leading causes of death and disability throughout the world.[
ML algorithms including RF, RR, and NB were all identified as effective prediction models for the unfavorable outcome or in-hospital mortality in TBI patients.[
The use of scoring algorithms SAPS II, APACHE II, and SOFA was found to have increased predictive power of inhospital mortality over LR, but no significant difference with overall 6-month mortality.[
The heterogeneity of input variables between ML models and studies limits the potential for cross-comparison. Those studies that had compatible methodologies were included in a small meta-analysis in an attempt to draw a quantitative conclusion regarding which model best predicts mortality. However, with the heterogeneity of input variables, inconsistency of outcome measurement, and variable criteria for TBI classification, this cannot be generalized to all TBI mortality predictions. A further prospective study with an increased sample size is necessary to definitively state, in which ML model is objectively most effective at mortality prediction. Furthermore, future studies should seek to standardize the necessary input variables for the operation of ML models. There is great inconsistency among the presented studies in the selection of input variables, with some studies only utilizing a few simple serum studies. While convenient for the provider, this limited input data may fail to capture a complete picture of the patient’s current condition. On the other hand, multiple studies employ a myriad of input variables including information that may not be easily accessible in an emergent situation, such as detailed radiological findings and hospital staffing statistics. [
CONCLUSION
TBI continues to be one of the leading causes of death and disability worldwide. This study reiterates the clinical utility of ML as an adjunct in patients with TBI. The use of ML to predict outcomes following TBI is entering clinical practice at an increasing rate and the present study reinforces the utility of these models. Using these models, simple admission data can be used to accurately predict the prognosis for individual patients. This can ultimately enhance the clinical decision-making process in terms of whether surgical intervention, medical management, or palliative care is most appropriate. There was a lack of consistency among the investigated studies with the selection of input variables used for predictive models; as a result, some models simply had more data to utilize for prediction, making inter-study comparisons more difficult. Further, research should utilize the core prediction variables identified in this review and apply these markers across a wide range of models and in multiple clinical settings. Given that the described models have demonstrated a robust ability to predict outcomes, there exists a significant degree of untapped potential in implementing ML to aid in neurosurgical decision-making. It is conceivable that these tools can be further expanded to guide and optimize patient treatment and perhaps alert neuro-care providers of patients at high risk of early neurological deterioration. Despite the increased use and predictive power of ML, it remains to be seen whether clinicians will routinely incorporate these models to guide clinical care following TBI.
Declaration of patient consent
Patient’s consent not required as there are no patients in this study.
Financial support and sponsorship
Nil.
Conflicts of interest
There are no conflicts of interest.
Disclaimer
The views and opinions expressed in this article are those of the authors and do not necessarily reflect the official policy or position of the Journal or its management. The information contained in this article should not be considered to be medical advice; patients should consult their own physicians for advice as to their specific medical needs.
References
1. Abujaber A, Fadlalla A, Gammoh D, Abdelrahman H, Mollazehi M, El-Menyar A. Prediction of in-hospital mortality in patients on mechanical ventilation post traumatic brain injury: Machine learning approach. BMC Med Inform Decis Mak. 2020. 20: 336
2. Abujaber A, Fadlalla A, Gammoh D, Abdelrahman H, Mollazehi M, El-Menyar A. Prediction of in-hospital mortality in patients with post traumatic brain injury using National Trauma Registry and Machine Learning Approach. Scand J Trauma Resusc Emerg Med. 2020. 28: 44
3. Adams R, Henry KE, Sridharan A, Soleimani H, Zhan A, Rawat N. Prospective, multi-site study of patient outcomes after implementation of the TREWS machine learning-based early warning system for sepsis. Nat Med. 2022. 28: 1455-60
4. Amorim RL, Oliveira LM, Malbouisson LM, Nagumo MM, Simoes M, Miranda L. Prediction of early TBI mortality using a machine learning approach in a LMIC population. Front Neurol. 2020. 10: 1366
5. Andelic N, Howe EI, Hellstrøm T, Sanchez MF, Lu J, Løvstad M. Disability and quality of life 20 years after traumatic brain injury. Brain Behav. 2018. 8: e01018
6. Areas FZ, Schwarzbold ML, Diaz AP, Rodrigues IK, Sousa DS, Ferreira CL. Predictors of hospital mortality and the related burden of disease in severe traumatic brain injury: A prospective multicentric study in Brazil. Front Neurol. 2019. 10: 432
7. Bonds BW, Yang S, Hu PF, Kalpakis K, Stansbury LG, Scalea TM. Predicting secondary insults after severe traumatic brain injury. J Trauma Acute Care Surg. 2015. 79: 85-90
8. Carlsen RW, Daphalapurkar NP. The importance of structural anisotropy in computational models of traumatic brain injury. Front Neurol. 2015. 6: 28
9. Churpek MM, Yuen TC, Winslow C, Meltzer DO, Kattan MW, Edelson DP. Multicenter comparison of machine learning methods and conventional regression for predicting clinical deterioration on the wards. Crit Care Med. 2016. 44: 368-74
10. Dewan MC, Rattani A, Gupta S, Baticulon RE, Hung YC, Punchak M. Estimating the global incidence of traumatic brain injury. J Neurosurg. 2019. 130: 1080-97
11. Dikmen SS, Machamer JE, Powell JM, Temkin NR. Outcome 3 to 5 years after moderate to severe traumatic brain injury 11 No commercial party having a direct financial interest in the results of the research supporting this article has or will confer a benefit upon the author(s) or upon any organization with which the author(s) is/are associated. Arch Phys Med Rehabil. 2003. 84: 1449-57
12. Fartoumi S, Emeriaud G, Roumeliotis N, Brossier D, Sawan M. Computerized decision support system for traumatic brain injury management. J Pediatr Intensive Care. 2015. 5: 101-7
13. Faul M, Coronado V, editors. Epidemiology of traumatic brain injury. Handbook of Clinical Neurology. USA: Elsevier; 2015. 127: 3-13
14. Feng J, Wang Y, Peng J, Sun M, Zeng J, Jiang H. Comparison between logistic regression and machine learning algorithms on survival prediction of traumatic brain injuries. J Crit Care. 2019. 54: 110-6
15. Gravesteijn BY, Nieboer D, Ercole A, Lingsma HF, Nelson D, van Calster B. Machine learning algorithms performed no better than regression models for prognostication in traumatic brain injury. J Clin Epidemiol. 2020. 122: 95-107
16. Hale AT, Stonko DP, Brown A, Lim J, Voce DJ, Gannon SR. Machine-learning analysis outperforms conventional statistical models and CT classification systems in predicting 6-month outcomes in pediatric patients sustaining traumatic brain injury. Neurosurg Focus. 2018. 45: E2
17. Heo J, Yoon JG, Park H, Kim YD, Nam HS, Heo JH. Machine learning-based model for prediction of outcomes in acute stroke. Stroke. 2019. 50: 1263-5
18. Herasevich V, Tsapenko M, Kojicic M, Ahmed A, Kashyap R, Venkata C. Limiting ventilator-induced lung injury through individual electronic medical record surveillance. Crit Care Med. 2011. 39: 34-9
19. Hyder AA, Wunderlich CA, Puvanachandra P, Gururaj G, Kobusingye OC. The impact of traumatic brain injuries: A global perspective. NeuroRehabilitation. 2007. 22: 341-53
20. Hyland SL, Faltys M, Hüser M, Lyu X, Gumbsch T, Esteban C. Early prediction of circulatory failure in the intensive care unit using machine learning. Nat Med. 2020. 26: 364-73
21. Kayhanian S, Young AM, Mangla C, Jalloh I, Fernandes HM, Garnett MR. Modelling outcomes after paediatric brain injury with admission laboratory values: A machine-learning approach. Pediatr Res. 2019. 86: 641-5
22. Kim KW, Lee J, Choi SH, Huh J, Park SH. Systematic review and meta-analysis of studies evaluating diagnostic test accuracy: A practical review for clinical researchers-part I. General guidance and tips. Korean J Radiol. 2015. 16: 1175-87
23. Lee J, Kim KW, Choi SH, Huh J, Park SH. Systematic review and meta-analysis of studies evaluating diagnostic test accuracy: A practical review for clinical researchers-part II. Statistical methods of meta-analysis. Korean J Radiol. 2015. 16: 1188-96
24. Maas AI, Stocchetti N, Bullock R. Moderate and severe traumatic brain injury in adults. Lancet Neurol. 2008. 7: 728-41
25. Majdan M, Plancikova D, Brazinova A, Rusnak M, Nieboer D, Feigin V. Epidemiology of traumatic brain injuries in Europe: A cross-sectional analysis. Lancet Public Health. 2016. 1: e76-83
26. Martins ET, Linhares MN, Sousa DS, Schroeder HK, Meinerz J, Rigo LA. Mortality in severe traumatic brain injury: A multivariated analysis of 748 Brazilian patients from Florianópolis city. J Trauma. 2009. 67: 85-90
27. Matsuo K, Aihara H, Nakai T, Morishita A, Tohma Y, Kohmura E. Machine learning to predict in-hospital morbidity and mortality after traumatic brain injury. J Neurotrauma. 2020. 37: 202-10
28. Moher D, Liberati A, Tetzlaff J, Altman DG. Preferred reporting items for systematic reviews and meta-analyses: The PRISMA Statement. Open Med. 2009. 3: e123-30
29. Muballe KD, Sewani-Rusike CR, Longo-Mbenza B, Iputo J. Predictors of recovery in moderate to severe traumatic brain injury. J Neurosurg. 2019. 131: 1648-57
30. Olsen CR, Mentz RJ, Anstrom KJ, Page D, Patel PA. Clinical applications of machine learning in the diagnosis, classification, and prediction of heart failure. Am Heart J. 2020. 229: 1-17
31. Pearn ML, Niesman IR, Egawa J, Sawada A, AlmenarQueralt A, Shah SB. Pathophysiology associated with traumatic brain injury: Current treatments and potential novel therapeutics. Cell Mol Neurobiol. 2017. 37: 571-85
32. PRISMA-P Group, Moher D, Shamseer L, Clarke M, Ghersi D, Liberati A. Preferred reporting items for systematic review and meta-analysis protocols (PRISMA-P) 2015 statement. Syst Rev. 2015. 4: 1
33. Rabinowitz AR, Levin HS. Cognitive sequelae of traumatic brain injury. Psychiatr Clin North Am. 2014. 37: 1-11
34. Raj R, Luostarinen T, Pursiainen E, Posti JP, Takala RS, Bendel S. Machine learning-based dynamic mortality prediction after traumatic brain injury. Sci Rep. 2019. 9: 17672
35. Raj R, Skrifvars M, Bendel S, Selander T, Kivisaari R, Siironen J. Predicting six-month mortality of patients with traumatic brain injury: Usefulness of common intensive care severity scores. Crit Care. 2014. 18: R60
36. Rau CS, Kuo PJ, Chien PC, Huang CY, Hsieh HY, Hsieh CH. Mortality prediction in patients with isolated moderate and severe traumatic brain injury using machine learning models. PLoS One. 2018. 13: e0207192
37. Rizoli S, Petersen A, Bulger E, Coimbra R, Kerby JD, Minei J. Early prediction of outcome after severe traumatic brain injury: A simple and practical model. BMC Emerg Med. 2016. 16: 32
38. Rusnak M. Giving voice to a silent epidemic. Nat Rev Neurol. 2013. 9: 186-7
39. Sawyer AM, Deal EN, Labelle AJ, Witt C, Thiel SW, Heard K. Implementation of a real-time computerized sepsis alert in nonintensive care unit patients. Crit Care Med. 2011. 39: 469-73
40. Senders JT, Arnaout O, Karhade AV, Dasenbrock HH, Gormley WB, Broekman ML. Natural and artificial intelligence in neurosurgery: A systematic review. Neurosurgery. 2018. 83: 181-92
41. Senders JT, Staples PC, Karhade AV, Zaki MM, Gormley WB, Broekman ML. Machine learning and neurosurgical outcome prediction: A systematic review. World Neurosurg. 2018. 109: 476-86.e1
42. Senders JT, Zaki MM, Karhade AV, Chang B, Gormley WB, Broekman ML. An introduction and overview of machine learning in neurosurgical care. Acta Neurochir. 2018. 160: 29-38
43. Shi HY, Hwang SL, Lee KT, Lin CL. In-hospital mortality after traumatic brain injury surgery: A nationwide population-based comparison of mortality predictors used in artificial neural network and logistic regression models. J Neurosurg. 2013. 118: 746-52
44. Stone JR, Wilde EA, Taylor BA, Tate DF, Levin H, Bigler ED. Supervised learning technique for the automated identification of white matter hyperintensities in traumatic brain injury. Brain Inj. 2016. 30: 1458-68
45. Donald R, Howells T, Piper I, Enblad P, Nilsson P. Forewarning of hypotensive events using a Bayesian artificial neural network in neurocritical care. J Clin Monit Comput. 2019. 33: 39-51
46. Whiting PF. QUADAS-2: A revised tool for the quality assessment of diagnostic accuracy studies. Ann Intern Med. 2011. 155: 529
47. Williams OH, Tallantyre EC, Robertson NP. Traumatic brain injury: Pathophysiology, clinical outcome and treatment. J Neurol. 2015. 262: 1394-6
48. Yarkoni T, Westfall J. Choosing prediction over explanation in psychology: Lessons from machine learning. Perspect Psychol Sci. 2017. 12: 1100-22
49. Zelnick LR, Morrison LJ, Devlin SM, Bulger EM, Brasel KJ, Sheehan K. Addressing the challenges of obtaining functional outcomes in traumatic brain injury research: Missing data patterns, timing of follow-up, and three prognostic models. J Neurotrauma. 2014. 31: 1029-38